(Please note that starting with perClass 3.x (June 2011), internal parameters of pipelines are not directly accessible)
Problem: Can I use PRTools with PRSD Studio?
Solution: Yes, PRTools users may benefit from many seamlessly integrated PRSD Studio tools.
PRTools is an academic toolbox developed in PRLab at TU Delft. Although PRSD Studio is a self-contained software without external dependencies, it provides close integration with PRTools enabling you to:
- Use powerful interactive tools on PRTools datasets
- Train scalable PRSD Studio classifiers on PRTools datasets
- Optimize PRTools classifiers with two- or multi-class ROC analysis
- Quickly execute many PRTools classifiers outside Matlab
In PRTools, data is stored in dataset objects. Although PRSD Studio directly accepts PRTools dataset objects in most situations, the two object formats may be easily converted if needed.
To convert PRTools dataset into sddata object use sddata
function:
>> a=gendatb
Banana Set, 100 by 2 dataset with 2 classes: [50 50]
>> b=sddata(a)
'Banana Set' 100 by 2 sddata, 2 classes: '1'(50) '2'(50)
Note that while PRTools supports numerical class names, PRSD Studio data
sets always contain string class names. This is to avoid confusion between
class indices and names. Only the default PRTools label set is converted
into sddata object. If additional sets of labels are stored in PRTools
dataset (either as labels or as custom targets), these need to be
transferred manually.
To convert PRSD Studio data set object into PRTools format, use dataset function:
>> load medical
>> a
'medical D/ND' 6400 by 11 sddata, 3 classes: 'disease'(1495) 'no-disease'(4267) 'noise'(638)
>> b=dataset(a)
6400 by 11 dataset with 3 classes: [1495 4267 638]
19.1. Leverage PRSD Studio interactive tools on PRTools datasets ↩
PRSD Studio interactive scatter may be used directly on PRTools dataset objects:
>> a=gendath
Highleyman Dataset, 100 by 2 dataset with 2 classes: [50 50]
>> sdscatter(a)
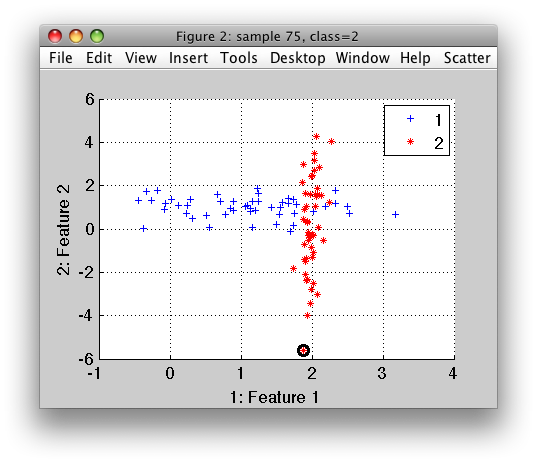
The scatter plot provides all functionality described in Chapter 6 including:
- hand-painting labels
- adding new label sets
- visualizing class overlap with per-feature distributions
For more information, see short videos.
Class overlap may be easily inspected using interactive feature
distribution plot sdfeatplot:
>> d
884 by 11 dataset with 3 classes: [140 194 550]
>> sdfeatplot(d)
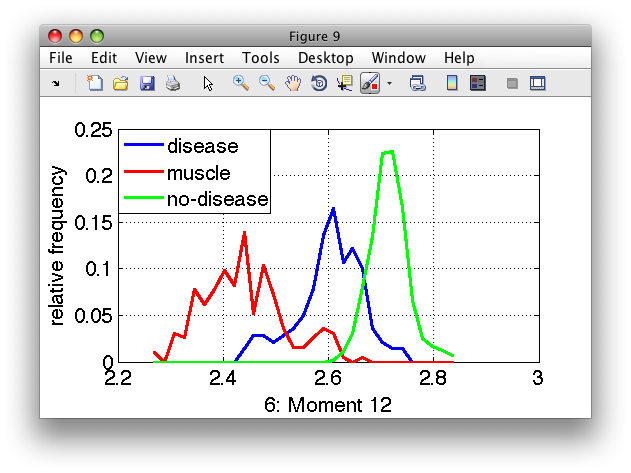
Use up/down cursor keys to switch between features.
19.2. Train fast and scalable classifiers ↩
With PRSD Studio you may always choose which classifier best suits your needs. You may seamlessly switch between training PRTools classifiers on PRSD Studio data sets and leveraging PRSD Studio classifiers without leaving PRTools.
PRSD Studio provides a number of highly scalable algorithms that yield both
high-speed training and execution on very large datasets. Lets take an
example of k-means clustering on a data set with 180 000 samples. We will
use the PRSD Studio command sdkmeans and ask for 100 clusters:
>> d
180000 by 2 dataset with 1 class: [180000]
>> tic; p=sdkmeans(d,100,'cluster'); toc
[class 'unknown' 100 clusters]
Elapsed time is 3.283098 seconds.
The clustering took 3.3 seconds. The result is a trained model p we may
apply to any data to get cluster labels.
>> tic; lab=d*sddecide(p); toc
Elapsed time is 0.346232 seconds.
For comparison, PRTools kmeans clustering of the same dataset took 26 minutes (PRSD Studio gives 440-times speedup):
>> tic; lab=kmeans(d,100); toc
Elapsed time is 1589.395077 seconds.
sdkmeans may be also used for multi-class data sets where it derives
k-means prototypes for k-NN classifier. In this way, we may very quickly
train highly scalable and robust non-parametric classifiers as illustrated in this
video.
19.2.1. Training PRSD Studio classifiers on PRTools datasets ↩
Any PRSD Studio classifiers may be trained on PRTools dataset object
directly, without conversion to sddata.
In this example, we train the Gaussian mixture model automatically estimating number
of mixture components from the data using PRSD Studio sdmixture command:
>> a
Highleyman Dataset, 2000 by 2 dataset with 2 classes: [1004 996]
>> p=sdmixture(a)
[class '1' initialization: 1 cluster EM:.............................. 1 comp]
[class '2' initialization: 5 clusters EM:.............................. 5 comp]
Mixture of Gaussians pipeline 2x2 2 classes, 6 components (sdp_normal)
Alternative way is to multiply the dataset with an untrained pipeline object. In this example, we scale data by standardization and then train feed-forward neural network on PRTools dataset:
>> pu=sdscale*sdneural
untrained pipeline 2 steps: sdscale+sdneural
>> p=a*pu
epochs (*100):..........
sequential pipeline 2x2 'standardization+Neural network'
1 standardization 2x2 (sdp_affine)
2 Neural network 2x2 (sdp_neural)
Note that to run this example, you don't need Neural Network Toolbox installed.
19.2.2. Training PRTools classifiers on PRSD Studio data sets ↩
Any PRTools classifier may be trained directly on PRSD Studio
sddata object by multiplication with the respective PRTools
untrained mapping.
In this example, we train PRTools Fisher linear discriminant directly on PRSD Studio medical data set:
>> b
'medical D/ND' 884 by 11 sddata, 3 'tissue' groups: 'disease'(140) 'no-disease'(550) 'muscle'(194)
>> w=b*fisherc
Fisher, 11 to 3 trained mapping --> stacked
We may visualize the classifier decisions in multi-dimensional space using:
>> sdscatter(b,sddecide(w))
Note that invoking fisherc on sddata object b using standard
function call results in error because PRTools does not recognize
sddata object.
>> w=fisherc(b)
{??? Error using ==> isdataset at 24
Dataset expected.
Error in ==> isvaldfile at 32
isdataset(a);
Error in ==> fisherc at 60
isvaldfile(a,1,2); % at least 1 object per class, 2 classes
}
To use this way of training, convert the sddata object into PRTools dataset:
>> w=fisherc(dataset(b))
Fisher, 11 to 2 trained classifier --> affine
19.3. Optimize PRTools classifiers with ROC analysis ↩
Performance of any PRTools classifier may be directly optimized using a comprehensive set of PRSD Studio ROC analysis tools.
This includes:
- ROC analysis for any two- or multi-class classifier
- converting any generative PRTools model into a detector (gaussm, parzenm)
- adding reject option to any PRTools classifier
Example 1: Multi-class ROC analysis of a PRTools classifier
In this example, we perform multi-class ROC analysis for linear discriminant on 8-class problem:
>> a=gendatm(10000)
Multi-Class Problem, 10000 by 2 dataset with 8 classes: [1255 1230 1296 1211 1246 1263 1260 1239]
>> [tr,ts]=gendat(a,0.5)
Multi-Class Problem, 5002 by 2 dataset with 8 classes: [628 615 648 606 623 632 630 620]
Multi-Class Problem, 4998 by 2 dataset with 8 classes: [627 615 648 605 623 631 630 619]
>> w=ldc(tr)
PR_Warning: getprior: ldc: No priors found in dataset, class frequencies are used instead
Bayes-Normal-1, 2 to 8 trained mapping --> normal_map
Soft outputs of the trained classifier:
>> out=ts*w
Multi-Class Problem, 4998 by 8 dataset with 8 classes: [627 615 648 605 623 631 630 619]
ROC analysis using sdroc command:
>> tic; r=sdroc(out), toc
..........
ROC (2000 w-based op.points, 9 measures), curop: 1827
est: 1:err(a)=0.21, 2:err(b)=0.24, 3:err(c)=0.00, 4:err(d)=1.00, 5:err(e)=0.03, 6:err(f)=0.34, 7:err(g)=0.19, 8:err
Elapsed time is 3.885148 seconds.
We may visualize the ROC surface using sddrawroc command:
>> sddrawroc(r);
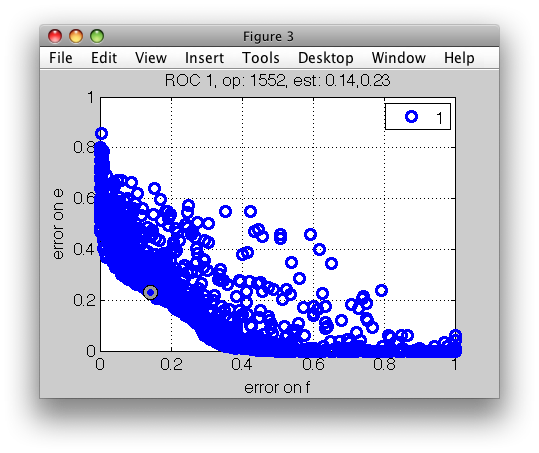
We may then use a number of methods to select desired operating point
including application of constraints or cost-sensitive optimization. For
example, we may require that the error on class f is smaller than 20%:
>> r2=constrain(r,'err(f)',0.2)
ROC (705 w-based op.points, 9 measures), curop: 1
est: 1:err(a)=0.27, 2:err(b)=0.18, 3:err(c)=0.00, 4:err(d)=1.00, 5:err(e)=0.20, 6:err(f)=0.19, 7:err(g)=0.20, 8:err
Once ready, we may get decisions at this operating point for new examples:
>> newdata=[1 0; -10 10]
newdata =
1 0
-10 10
>> dec=sddata(newdata)*w*r2
sdlab with 2 entries, 2 groups: 'a'(1) 'h'(1)
19.4. Quickly deploy classifiers out of Matlab ↩
PRSD Studio offers an easy-to-use set of tools for classifier deployment in custom applications. Main benefits are:
- Quickly embed classifiers trained in PRTools into custom applications
- Significantly speed up your classifier execution
Any supported trained PRTools mapping may be converted into PRSD Studio
pipeline using sdconvert command:
>> b
200 by 2 dataset with 2 classes: [100 100]
>> w=fisherc(b)
Fisher, 2 to 2 trained classifier --> affine
>> p=sdconvert(w)
sequential pipeline 2x2 'Fisher'
1 Fisher 2x2 (sdp_affine)
2 Sigmoid 2x2 (sdp_sigmoid)
Pipeline objects may be exported for execution outside Matlab using the
sdexport command. For information on classifier deployment, see the Chapter 14.
One-step conversion of a PRTools mapping into pipeline and adding an
operating point is possible using the sddecide function. Below, you can
see an example of exporting AdaBoost classifier, trained in PRTools, for
execution outside Matlab.
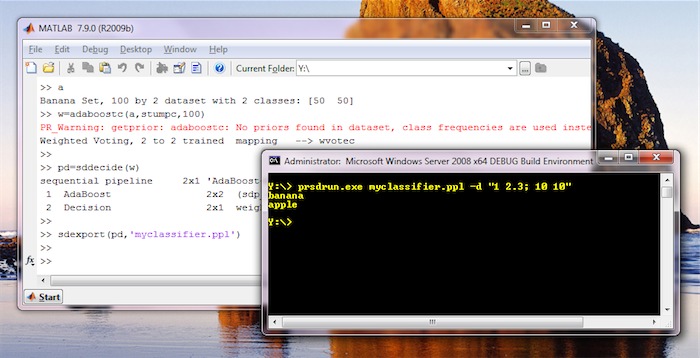
19.4.1. Supported PRTools mappings ↩
The PRSD deployment technology seamlessly supports more than 40 most common PRTools classifiers. Also any sequential, stacked (same feature space) or parallel (different feature spaces) combination is supported.
adaboostc(AdaBoost) - only with decision tree base learner (treecorstumpc)baggingc(Bagging)bhatm(Bhattacharryya linear feature extraction)chernoffm(Chernoff mapping: Suboptimal discrimination linear mapping)emclust(Expectation-Maximization clustering) - any supported modelfdsc(Feature based Dissimilarity Space Classification)featsel(Feature selection mapping)fisherc(Fisher's Least Square Linear Classifier)fisherm(Fisher projection, LDA)gaussm(Mixture of Gaussians density estimate)klldc(Linear classifier built on the KL expansion of the common covariance matrix)klm(Karhunen-Loeve Mapping)klms(Karhunen-Loeve Mapping, followed by scaling)knnc(K-Nearest Neighbor Classifier)ldc(Linear Bayes Normal Classifier)loglc(Logistic Linear Classifier)maxc(Maximum classifier combiner)meanc(Mean classifier combiner)minc(Min classifier combiner)mogc(Mixture of Gaussian classifier)nlfisherm(Non-linear Fisher Mapping according to Marco Loog)nmc(Nearest mean classifier)nmsc(Nearest Mean Scaled Classifier)parallel(Parallel classifier combination)parzenc(Parzen density classifier)parzenm(Parzen density estimator) - only scalar smoothingpca(Principal Component Analysis)pcaklm(Principal Component Analysis/Karhunen-Loeve Mapping)pcldc(Linear classifier using PC expansion on the joint data)perlc(Linear perceptron classifier)prodc(Product classifier combiner)proxm(Proximity mapping) - Distance, RBF, Polynomialqdc(Quadratic Bayes Normal Classifier)scalem(Scaling projection)sequential(Sequence of mappings)sigm(Sigmoid map)stacked(Stacked combination of classifiers)stumpc(Decision stump)svc(Support vector machine) - Linear, RBF, Polynomial kerneltreec(Decision tree)udc(Uncorrelated normal based quadratic Bayes classifier)weakc(Weak classifier) - any supported model
