This chapter describes methods to reduce number of feature by extraction or selection.
- 12.1. Feature extraction
- 12.1.1. Principal Component Analysis (PCA)
- 12.1.2. Linear Discriminant Analysis (LDA)
- 12.2. Feature space expansion
- 12.3. Feature selection
- 12.3.1. Manual feature selection
- 12.3.2. Individual feature selection (feature ranking)
- 12.3.3. Random feature selection
- 12.3.4. Forward search
- 12.3.5. Backward search
- 12.3.6. Floating search
- 12.3.7. Initialization of the selection searches
- 12.3.8. Using a decision tree classifier for feature selection
An important pre-requisite for training robust classifiers is construction of an informative data representation. This chapter is organized into two parts. First, we discuss feature extraction and then feature selection methods.
Two basic types of dimensionality reduction include feature extraction and feature selection. Feature extraction results in a lower-dimensional space constructed using information from all original features. Feature selection, on the other hand, chooses a smaller subset of the original features. This is useful to identify the informative features, and to limit computational demands when executing the recognition system on new observations.
12.1. Feature extraction ↩
Two prominent feature extraction methods are Principal Component Analysis (PCA) and Linear Discriminant Analysis (LDA). The major difference is that PCA is an unsupervised approach (not using ground-truth labels) while LDA is fully supervised.
12.1.1. Principal Component Analysis (PCA) ↩
PCA derives lower-dimensional linear subspace by preserving a specific
fraction of variation in the original data. It is implemented using the
sdpca command. This feature extraction method ia unsupervised. This means
that the class labels are not taken into account. Therefore, the presence
of labels in the data set does not alter the resulting PCA projection.
Running sdpca without additional arguments results in a decorrelation of
the data (mere rotation) preserving all information in the original data
set. This is useful, for example, when building classifiers that makes use
of thresholding (decision trees) or assumptions of independence (naive
Bayes).
>> p=sdpca(a)
PCA pipeline 300x20 100% of variance (sdp_affine)
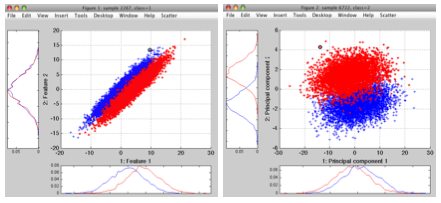
In order to reduce number of features, we may either provide the desired dimensionality directly or specify the fraction of preserved variance:
>> p=sdpca(a,3) % specifying output dimensionality
PCA pipeline 300x3 74% of variance (sdp_affine)
>> p=sdpca(a,0.95) % specifying fraction of preserved variance
PCA pipeline 300x16 95% of variance (sdp_affine)
Note that sdpca always displays how much of original data variance is
preserved.
Often, we build PCA subspace to train a classifier. sdpca allows us to
quickly select the dimensionality minimizing error of a specified
classifier:
>> p=sdpca(a,sdlinear)
.......... 48D subspace (frac=0.40), err=0.05
Affine pipeline 300x48 (sdp_affine)
The sdpca splits the data set a into training and validation subsets
(by default 50/50). The model (in our example sdlinear) is trained on the
training part and its error is evaluated on the validation part. The
resulting PCA projection uses dimensionality minimizing the error. We may
adjust the fraction of validation data using 'tsfrac' option.
As with any other perClass routine that performs internal splitting of data, we may split externally avoiding otherwise present variability:
>> [tr,val]=randsubset(a,0.8)
>> p=sdpca(tr,sdlinear,'test',val)
........ 67D subspace (frac=0.40), err=0.06
Affine pipeline 300x67 (sdp_affine)
TIP: It is a good practice to perform PCA on a data set we start working on in order to understand its complexity. If the PCA preserving some 95 or 99% of variance results in significant dimensionality reduction, we probably deal with data located along a simple manifold structure. This is often happening for signals or image data sets. If the resulting dimensionality is high compared with the original number of features, we face a complicated problem and may need more complex classifier and consequently larger number of training examples.
12.1.2. Linear Discriminant Analysis (LDA) ↩
LDA is a supervised feature extraction method that finds a linear subspace maximizing separability between classes. The dimensionality of the resulting subspace is fixed to the minimum between: number of features, number of samples, number of classes - 1). Usually, the output dimensionality is determined by the number of classes. For a two-class problem LDA results in a projection on a line, for thee-class problem on a 2D plane.
>> a
'medical D/ND' 6400 by 11 sddata, 3 classes: 'disease'(1495) 'no-disease'(4267) 'noise'(638)
>> p=sdlda(a)
LDA pipeline 11x2 (sdp_affine)
LDA projection may be computed for data sets with number of features larger than number of classes by using the pseudo-inverse. However, we should carefuly investigated the generalization capability of such projections, i.e. the capability of reliably mapping unseen data.
One possible remedy for poor generalization capability of LDA in high-dimensional small-sample size problems is to first perform PCA and estimate LDA projection only in lower-dimensional subspace of the original data set.
>> d
'Difficult Dataset' 100 by 200 sddata, 2 classes: '1'(48) '2'(52)
>> p=sdpca([],10)*sdlda
untrained pipeline 2 steps: sdpca+sdlda
>> p2=d*p
sequential pipeline 200x1 'PCA+LDA'
1 PCA 200x10 38%% of variance (sdp_affine)
2 LDA 10x1 (sdp_affine)
12.2. Feature space expansion ↩
Special form of representation building is feature space expansion. Instead of reducing the dimensionality, it generates new features based on the original ones (therefore it's similar to extraction).
sdexpand implements polynomial expansion adding powers or cross-products
of existing features. This is useful in order to build non-linear or more
complex classifiers using simpler models. It is used by default in the logistic classifier
sdlogistic.
>> a
'Fruit set' 260 by 2 sddata, 3 classes: 'apple'(100) 'banana'(100) 'stone'(60)
>> ppoly=sdexpand(a,5)
Feature expansion pipeline 2x11
>> ppoly.lab'
1 F1:1
2 F2:2
3 F1^2
4 F2^2
5 F1^3
6 F2^3
7 F1^4
8 F2^4
9 F1^5
10 F2^5
11 F1*F2
>> b=a*ppoly
'Fruit set' 260 by 11 sddata, 3 classes: 'apple'(100) 'banana'(100) 'stone'(60)
12.3. Feature selection ↩
Feature selection strategies aim at selecting an informative subset of
features out of the complete set. The goodness of a solution is judged
based on a criterion or on the classification error. perClass includes fast
feature selection via the sdfeatsel function. By default, sdfeatsel
selects features by minimizing the error of 1-NN classifier trained on a
part of the data and tested on the remaining test set. Leveraging
classifier error as a feature selection criterion has the advantage of
providing a clear decision on how many features are needed.
12.3.1. Manual feature selection ↩
In the simplest scenario we know what features we want to select. We may provide the data set and desired feature indices or names manually.
This is useful, for example, when we build a cascaded system with several classifiers, each trained on a different subset of the entire feature set.
Defining the subset manually by input feature indices:
>> pf=sdfeatsel(a,[1:4 8])
Feature subset pipeline 11x5
We can see the details of the selection pipeline with the transpose operator:
>> pf'
Feature subset pipeline 11x5
1 Feature subset 11x5
inlab: 'StdDev','Skew','Kurtosis','Energy Band 1','Energy Band 2',...
lab: 'StdDev','Skew','Kurtosis','Energy Band 1','Moment 02'
output: data
ind: feature subset indices
The pf.lab field contains the feature labels of the selected features
>> pf.lab
sdlab with 5 entries:
'StdDev','Skew','Kurtosis','Energy Band 1','Moment 02'
The pf.inlab stores the input feature names. This is useful later on when
we load a classifier trained earlier in our project. We can always see what
features the pipeline uses from the input data.
To select a subset of features, simply apply the feature seletion pipeline to the data set:
>> a2=a*pf
'medical D/ND' 5762 by 5 sddata, 2 classes: 'disease'(1495) 'no-disease'(4267)
>> a2.featlab'
1 StdDev
2 Skew
3 Kurtosis
4 Energy Band 1
5 Moment 02
Feature subsets may be also specified by cell array of names of regular expressions:
>> pf=sdfeatsel(a,{'Skew','Moment 02'})
Feature subset pipeline 11x2
>> pf=sdfeatsel(a,'/Moment|Kurt')
Feature subset pipeline 11x5
>> pf.lab
sdlab with 5 entries:
'Kurtosis','Moment 12','Moment 20','Moment 02','Moment 13'
12.3.2. Individual feature selection (feature ranking) ↩
The individual search evaluates each feature separately. The advantage of individual search is high speed. It is therefore useful for pre-selection of a candidate feature subset from a large set of features. Note, however, that individually poor features may yield high class separability when used together.
>> pf=sdfeatsel(a,'individual')
individual:
Feature subset pipeline 11x1
>> pf.lab
sdlab with one entry: 'StdDev'
Because of internal splitting into a subset used for classifier training and subset for criteria evaluation, this default selection process may yield different result each run.
>> pf=sdfeatsel(a,'individual')
individual:
Feature subset pipeline 11x1
>> pf.lab
sdlab with one entry: 'Skew'
To make selection repeatable, you may split the data set yourself outside
and provide the two sets separately using the 'test' option. The data set
tr is then used for classifier training and the test set ts for error
estimation.
>> [tr,ts]=randsubset(a,0.5)
'medical D/ND' 3199 by 11 sddata, 3 classes: 'disease'(747) 'no-disease'(2133) 'noise'(319)
'medical D/ND' 3201 by 11 sddata, 3 classes: 'disease'(748) 'no-disease'(2134) 'noise'(319)
>> pf=sdfeatsel(tr,'test',ts,'individual')
individual:
Feature subset pipeline 11x1
To inspect the feature selection process sdfeatsel provides, as the
second output, the structure with detailed information.
>> [pf,res]=sdfeatsel(tr,'test',ts,'individual')
individual:
Feature subset pipeline 11x1
res =
feat: [6 8 2 9 7 1 11 5 3 10 4]
crit: [1x11 double]
best_feat: 6
best_crit: 0.5081
method: 'individual ranking'
12.3.3. Random feature selection ↩
Random search evaluates a large set of random feature subsets and returns the best one. It is useful to implement more complex feature selection strategies such as genetic feature selection or as an initialization of greedy searches. For example, the forward search suffers if all features judged individually are poor. Bad choices made early on in greedy search cannot be undone. Random search allows us to start the greedy search from a meaningful feature subset.
>> [pf,res]=sdfeatsel(a,'random');
>> pf
Feature subset pipeline 11x5
The res structure contains all analyzed feature subsets in a binary
matrix res.subsets (subsets vs features). The criterion values are stored
into res.crit.
>> res
subsets: [200x11 uint8]
crit: [200x1 double]
best_subset: [1 2 3 9 11]
best_count: 5
best_crit: 0.2868
method: 'random search'
The number of evaluated random subsets can be changed with 'count' option. The length of subset sizes can be controlled with 'bounds' option.
12.3.4. Forward search ↩
The forward search is a greedy strategy. It start with the best individual feature and adds to it the feature that together with the first provides the best result. The procedure is repeated until all features are used. Therefore, in the forward search, the merit of additional features are judged together with the already selected subset.
>> [pf,res]=sdfeatsel(tr,'test',ts,'forward');
forward:...........
Feature subset pipeline 11x7 (sdp_fsel)
res =
method: 'forward search'
improvement: 1
best_subset: [6 11 9 3 10 7 8]
best_count: 7
best_crit: 0.2699
feat_init: []
feat: [6 11 9 3 10 7 8 2 1 5 4]
crit: [1x11 double]
Number of features is selected automatically because the selection criteria uses classifier error. If no test set is provided the input data set is split randomly into two parts, one used to train 1-NN classifier used as criterion, and the other to estimate the performance. Due to the random splitting, repeating the search on the same data may result in different subset of selected features.
>> [pf,res]=sdfeatsel(a)
res =
method: 'forward search'
improvement: 1
best_subset: [6 11 8 3 7 1 9 2]
best_count: 8
best_crit: 0.2655
feat_init: []
feat: [6 11 8 3 7 1 9 2 10 5 4]
crit: [1x11 double]
>> [pf,res2]=sdfeatsel(a)
forward:...........
Feature subset pipeline 11x5 (sdp_fsel)
res2 =
method: 'forward search'
improvement: 1
best_subset: [6 11 8 3 7]
best_count: 5
best_crit: 0.2572
feat_init: []
feat: [6 11 8 3 7 9 10 2 1 5 4]
crit: [1x11 double]
The default splitting fraction is 0.75 (75% of data used for training). The
fraction may be changed using 'trfrac' option. Additional information on
the feature selection process may be retrieved using the second output: the
structure res provides the best subset found and also all subsets
analyzed together with the criteria values.
We may start the forward search from an initial subset using the 'from' option providing feature indices or selection pipeline:
>> [pf,res]=sdfeatsel(a,'from',[ 1 3 4])
forward:initial set: crit=0.240000
.....
Feature subset pipeline 8x7 Best forward subset (1-NN)
res =
method: 'forward search'
improvement: 1
best_subset: [1 3 4 7 6 8 5]
best_crit: 0.2000
best_count: 7
feat_init: [1 3 4]
crit_init: 0.2400
feat: [1 3 4 7 6 8 5 2]
crit: [NaN NaN 0.2400 0.2200 0.2200 0.2200 0.2000 0.2200]
The forward search may be limited (and sped up) using the 'steps' option. We may, for example let it perform only 5 steps:
>> [pf,res]=sdfeatsel(a,'steps',5)
forward:.....
Feature subset pipeline 8x1 Best forward subset (1-NN)
res =
method: 'forward search, making 5 steps, returning best subset'
improvement: 1
best_subset: 2
best_count: 1
best_crit: 0.2800
feat_init: []
feat: [2 7 4 1 5]
crit: [0.2800 0.2800 0.3200 0.3200 0.3000]
12.3.5. Backward search ↩
The backward search starts from all features and gradually removes the features until the performance improves.
>> [pf,res]=sdfeatsel(tr,'test',ts,'backward')
backward:..........
Feature subset pipeline 11x6 (sdp_fsel)
res =
method: 'backward search'
improvement: 1
best_subset: [1 11 9 3 2 8]
best_count: 6
best_crit: 0.2833
feat_init: []
feat: [1 11 9 3 2 8 7 6 10 5 4]
crit: [1x11 double]
Also the backward searches may be limited with the 'steps' option:
>> [pf,res]=sdfeatsel(a,'backward','steps',5)
backward:initial set: crit=0.260000
1:removing 4 crit=0.200000
2:removing 6 crit=0.200000
3:removing 7 crit=0.200000
4:removing 3 crit=0.220000
5:removing 2 crit=0.240000
Feature subset pipeline 8x5 Best backward subset (1-NN)
res =
method: 'backward search, making 5 steps, returning best subset'
improvement: 1
best_subset: [1 2 3 5 8]
best_count: 5
best_crit: 0.2000
feat_init: []
feat: [1 5 8 2 3 7 6 4]
crit: [NaN NaN 0.2400 0.2200 0.2000 0.2000 0.2000 0.2600]
12.3.6. Floating search ↩
The floating search combines multiple forward and backward steps gradually improving the best feature subset found. If not initial feature set is given, the floating search is initialized by the random search.
>> [pf,res]=sdfeatsel(tr,'test',ts,'floating')
floating: random:(25), backward:........(22), forward:.........(44), backward:...........(44), forward:........(44)
Feature subset pipeline 60x44
res =
best_subset: [1x41 double]
best_crit: 0.0191
feat: {1x5 cell}
feat_count: [25 22 44 41 44]
crit: [0.0765 0.0383 0.0281 0.0191 0.0281]
method: 'floating'
stop: 5
Floating search uses full forward and backward search steps. Therefore, it
cannot provide a single nested list of features. Instead, the res
structure contains a cell array with a feature subset for each floating
step and corresponding criterion value.
>> res.feat
ans =
Columns 1 through 4
[1x25 double] [1x22 double] [1x44 double] [1x41 double]
Column 5
[1x44 double]
The res structure also provides lengths of feature subsets in
res.feat_count field. Thereby, we may quickly visualize the process of
floating search minimizing the criterion:
>> figure; plot(res.feat_count,res.crit,'b-x')
>> xlabel('feature count')
>> ylabel('criterion (1-NN error) on the test set')
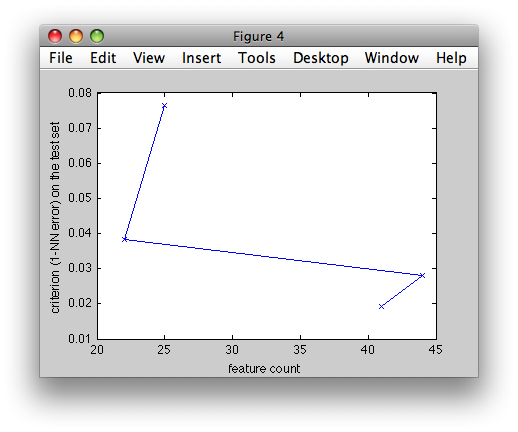
The x axis shows the number of features in each floating step, the y axis then the corresponding 1-NN error on the test set.
12.3.7. Initialization of the selection searches ↩
Forward, backward or floating search may be initiated from an existing subset of features. For example, we can start the forward search from the random search results:
>> [pf,res]=sdfeatsel(tr,'test',ts,'random');
>> [pf2,res2]=sdfeatsel(tr,'test',ts,'forward','from',pf);
forward:......
Feature subset pipeline 11x6 (sdp_fsel)
res2 =
method: 'forward search'
improvement: 1
best_subset: [1 2 3 9 11 8]
best_crit: 0.2833
best_count: 6
feat_init: [1 2 3 9 11]
crit_init: 0.2868
feat: [1 2 3 9 11 8 7 6 10 5 4]
crit: [0.2833 0.2837 0.2886 0.2985 0.3202 0.3755]
Additional considerations. In this example, we first run the forward search:
>> [pf1,res1]=sdfeatsel(tr,'test',ts)
forward:...........
Feature subset pipeline 11x7 (sdp_fsel)
res1 =
method: 'forward search'
improvement: 1
best_subset: [8 11 1 3 7 9 6]
best_count: 7
best_crit: 0.2961
feat_init: []
feat: [8 11 1 3 7 9 6 2 10 5 4]
crit: [1x11 double]
If we attempt to initiate another forward search from the subset found
(pf1), we will not improve:
>> [pf2,res2]=sdfeatsel(tr,'test',ts,'from',pf1)
forward:
Feature subset pipeline 11x7 (sdp_fsel)
res2 =
method: 'forward search'
improvement: 0
best_subset: [8 11 1 3 7 9 6]
best_crit: 0.2961
best_count: 7
feat_init: [8 11 1 3 7 9 6]
crit_init: 0.2961
feat: [8 11 1 3 7 9 6 2 10 5 4]
crit: [0.3160 0.3160 0.3527 0.3850]
However, if we perform a backward search we may reach better performance by removing some of the features:
>> [pf2,res2]=sdfeatsel(tr,'test',ts,'from',pf1,'backward')
backward:......
Feature subset pipeline 11x5 (sdp_fsel)
res2 =
method: 'backward search'
improvement: 1
best_subset: [7 11 8 3 6]
best_count: 5
best_crit: 0.2789
feat_init: [8 11 1 3 7 9 6]
feat: [7 11 8 3 6 9 1]
crit: [0.5377 0.4268 0.3429 0.2976 0.2789 0.2823 0.2961]
12.3.8. Using a decision tree classifier for feature selection ↩
Decision tree classifier may be used as a feature selection technique. For details, see this example.
